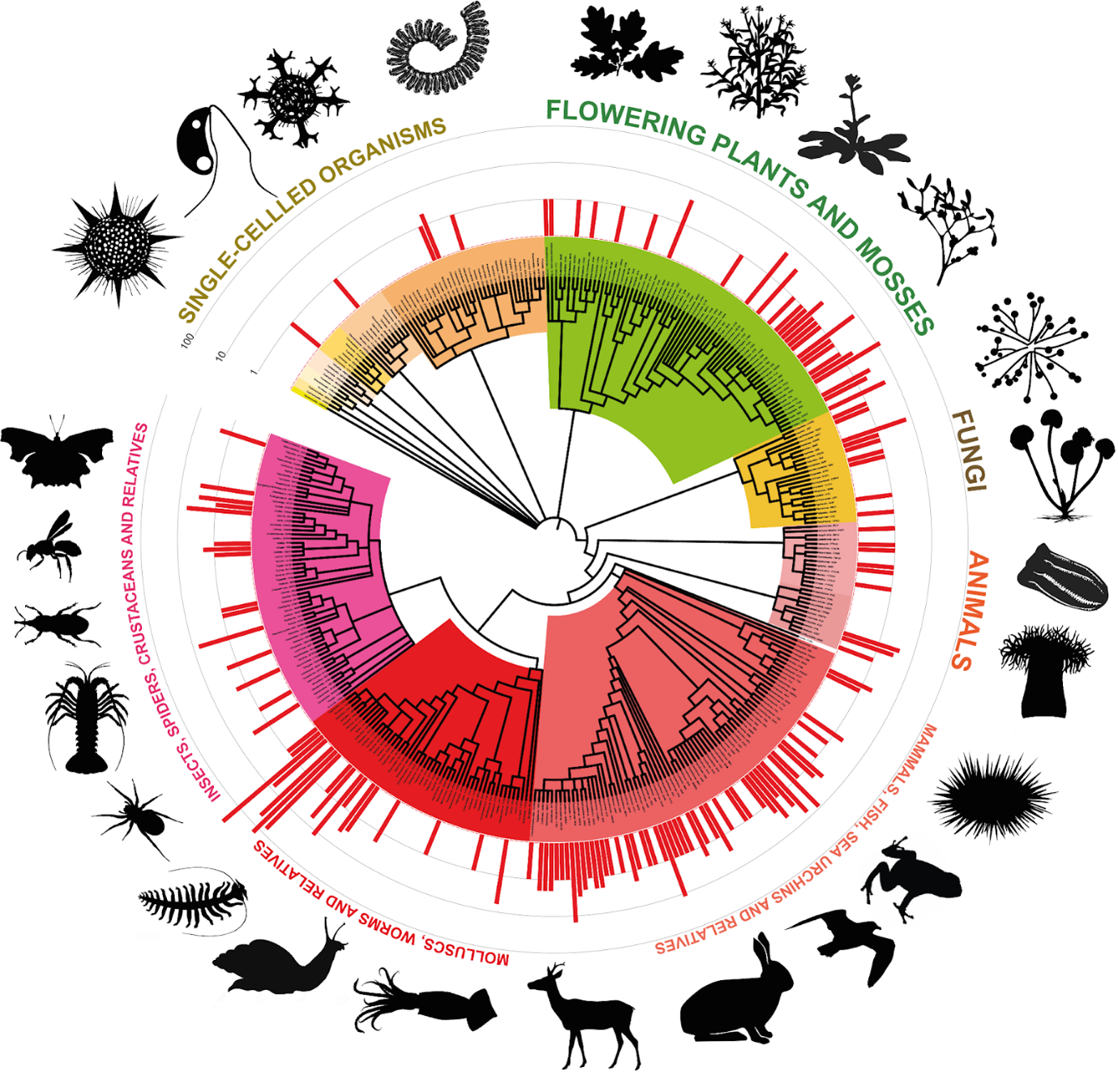
In an ambitious effort to map the biological blueprint of a region, researchers have launched the Darwin Tree of Life project, a massive scientific undertaking aimed to sequence the genomes of all complex life forms across the UK and Ireland. By digitizing the genetic codes of thousands of species, the initiative seeks to create a comprehensive genomic library that will fundamentally change how scientists track biodiversity and combat extinction.
The project represents a shift in biological research, moving from the study of individual species to a holistic, “large data” approach to ecology. For someone who spent years in software engineering before moving into tech reporting, the scale of this project is striking; it is essentially an attempt to build a searchable, high-resolution database of nature’s source code. The goal is to provide a definitive genetic record for every eukaryotic organism—ranging from fungi and mosses to complex animals—found within the British Isles.
By utilizing advanced next-generation sequencing (NGS) and bioinformatics, the project aims to fill critical gaps in the current global understanding of biodiversity. Even as many flagship species have had their genomes sequenced, the vast majority of “less charismatic” organisms—the insects, soil microbes, and obscure plants that sustain ecosystems—remain genomic mysteries. The Darwin Tree of Life project intends to rectify this by prioritizing these overlooked species.
The Mechanics of Genomic Mapping
At its core, the project is about more than just collecting DNA. It is about creating a reference framework that allows scientists to identify species more accurately and understand their evolutionary relationships. Traditional taxonomy, which relies on physical characteristics, can be misleading due to convergent evolution or the existence of “cryptic species”—organisms that look identical but are genetically distinct.
The research team employs a strategy of targeted sampling across diverse habitats. By sequencing the full genomes of these organisms, researchers can identify the specific genes responsible for climate resilience, disease resistance, and metabolic efficiency. This data is then uploaded to open-access repositories, ensuring that the global scientific community can leverage the findings for conservation and biotechnology.
The scale of the effort is managed through a collaborative network of universities and research institutions. The process typically involves several key stages:
- Field Sampling: Collecting biological specimens from varied ecological zones across the UK and Ireland.
- DNA Extraction: Isolating high-quality genetic material from the collected tissue.
- High-Throughput Sequencing: Using platforms like Pacific Biosciences (PacBio) or Oxford Nanopore to read long strands of DNA.
- Bioinformatic Assembly: Using computational algorithms to piece together the fragmented reads into a complete genome.
- Annotation: Identifying the functions of the genes within the sequenced genome.
Why a Comprehensive Genetic Library Matters
The implications of the Darwin Tree of Life project extend far beyond academic curiosity. In an era of rapid environmental decline, having a genetic “snapshot” of current biodiversity is a critical insurance policy. If a species goes extinct, its genetic information—and the biological secrets it holds—is lost forever unless it has been sequenced.
From a conservation standpoint, this data allows for “environmental DNA” (eDNA) monitoring. By sequencing a sample of water or soil, scientists can detect the presence of rare or invasive species without ever seeing them, simply by matching the DNA fragments found in the environment to the reference genomes created by the project. This allows for real-time tracking of species migration and the early detection of ecological threats.
the project has significant potential for the biotechnology and pharmaceutical industries. Many of the complex life forms being sequenced produce unique secondary metabolites—chemicals used for defense or communication—that could lead to the discovery of new antibiotics or sustainable materials. By mapping these genomes, researchers can identify the genetic pathways that produce these compounds, potentially allowing for synthetic production without needing to harvest rare wild organisms.
Project Scope and Strategic Focus
| Objective | Methodology | Expected Outcome |
|---|---|---|
| Biodiversity Baseline | Genome sequencing of UK/Ireland eukaryotes | Comprehensive genetic catalog of regional life |
| Species Identification | Comparative genomic analysis | Resolution of cryptic species and taxonomic errors |
| Conservation Tooling | eDNA reference library development | Non-invasive monitoring of endangered wildlife |
| Bioprospecting | Functional gene annotation | Discovery of novel biochemical properties |
Overcoming Computational and Biological Hurdles
Sequencing “all” complex life is a daunting task, not just biologically but computationally. Some plant genomes are gargantuan, containing far more DNA than a human genome, often filled with repetitive sequences that are notoriously difficult to assemble. This is where the intersection of biology and computer science becomes critical. The project relies on massive computing clusters and sophisticated algorithms to handle the petabytes of data generated.
We find also logistical challenges. Sampling across the diverse terrains of the UK and Ireland—from the Scottish Highlands to the bogs of Ireland—requires extensive field coordination. The project must balance the necessitate for comprehensive sampling with the ethical necessity of minimizing the impact on the extremely species it seeks to protect.
The project is aligned with broader international efforts, such as the Earth BioGenome Project, which shares the goal of sequencing all known eukaryotic species on Earth. By focusing on the UK and Ireland, the Darwin Tree of Life project provides a high-resolution regional model that can be replicated in other parts of the world.
The Road Ahead for UK Biodiversity
As the project progresses, the focus will shift from initial sequencing to the deeper analysis of how these genomes interact with changing environments. The next phase involves integrating genomic data with climatic and geographical data to predict how species will respond to rising temperatures and habitat fragmentation.
The project’s success will be measured by the growth of its open-access database and the number of species transitioned from “unknown” to “sequenced.” While the goal of sequencing every complex life form is an iterative process, the establishment of this genomic infrastructure provides a permanent resource for future generations of biologists and conservationists.
The project continues to expand its sampling efforts, with ongoing updates expected as new genomes are assembled and annotated in the public domain. For more information on genomic research and biodiversity, official updates are typically released through the participating university consortia and the Natural England framework.
This article is provided for informational purposes and does not constitute biological or environmental policy advice.
We would love to hear your thoughts on the intersection of big data and conservation. Please share this story or leave a comment below to join the conversation.