Inside the nucleus of every human cell, DNA does not sit as a loose string, but as a meticulously folded three-dimensional architecture. For years, biologists have grappled with a paradox: while the proteins that maintain this structure seem essential, removing them often leaves the cell’s overall gene activity surprisingly untouched. Although, new research suggests that this stability is a facade that hides a critical vulnerability in how we develop.
A study published April 13 in Nature Genetics reveals that disrupting genome architecture selectively impairs developmental genes, specifically those responsible for transforming a generic stem cell into a specialized organ cell. While most of the genome possesses a “molecular memory” that allows it to function despite structural collapse, the genes that dictate cell identity—such as those for the heart, brain, or liver—are far more fragile.
The findings, led by researchers at Weill Cornell Medicine, provide a potential explanation for why mutations in structural proteins lead to severe developmental disorders and cancers, even when the majority of the cell’s genetic machinery appears to be working normally.
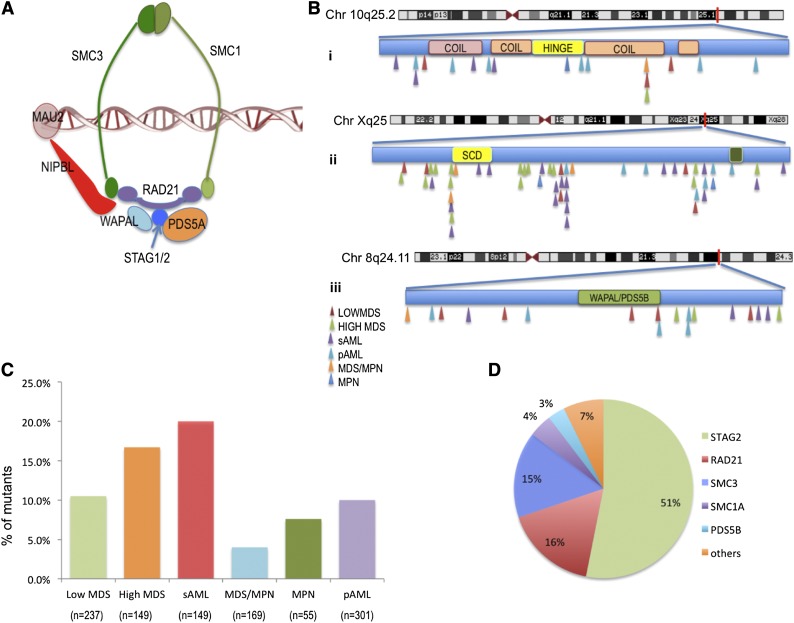
At the center of this process is a protein complex called cohesin. Cohesin acts as a molecular architect, looping DNA to bring distant regulatory elements into physical contact with the genes they control. This proximity is what allows a cell to “flip the switch” on specific genes, maintaining the cell’s identity and ensuring it performs its designated role in the body.
The Paradox of Genomic Resilience
The research team, led by senior author Effie Apostolou, an associate professor of molecular biology in medicine and member of the Sandra and Edward Meyer Cancer Center at Weill Cornell Medicine, sought to test the limits of this architecture. To do so, they focused on mouse embryonic stem cells—cells with the unique ability to become any tissue in the body.
The team targeted a specific, high-stress window: the moments immediately following cell division. This is the point where the entire genome must be rebuilt from scratch and the resulting daughter cells must “decide” whether to remain stem cells or differentiate into specialized types. By temporarily disabling cohesin at this exact juncture, the researchers could observe the immediate fallout of a collapsed genomic structure.
The results were unexpected. Using three-dimensional mapping techniques, the team confirmed that without cohesin, the DNA loops failed to re-form, and the overall genomic structure was severely disrupted. Yet, for the majority of genes, this structural chaos did not lead to functional failure.
“But then came the big surprise,” Apostolou said. “Most genes were largely unaffected.”
According to the study’s first author, graduate student UkJin Lee, this suggests the existence of a “resilient molecular memory” that persists through cell division. This memory allows the stem cell program to reactivate even when the primary architectural protein is missing, implying that other, yet-to-be-identified factors are helping the cell remember its identity.
Why Developmental Genes are Vulnerable
While the stem cells showed remarkable resilience, the stability vanished the moment the researchers “pushed” those cells to differentiate into specialized types. When the cells attempted to become specific tissues, a slight but vital group of genes failed to activate.
These “vulnerable genes” are typically those encoding transcription factors—the master regulators that direct cell identity. Unlike the resilient genes, these developmental regulators are often sequestered in isolated regions of the genome. They rely almost exclusively on cohesin to bridge the gap between the gene and the distant DNA elements that enhance their activity.
When this bridge is broken, the cell cannot trigger the necessary genetic programs. This failure can lead to a cascade of errors: genes that should be “on” remain “off,” or the wrong genes are activated entirely. This mechanism helps explain the pathology of cohesinopathies—a group of disorders characterized by physical and cognitive developmental impairments—as well as the prevalence of cohesin mutations in various cancers.
Impact of Cohesin Loss on Cell Function
| Gene Category | Structural Impact | Functional Outcome | Role in Body |
|---|---|---|---|
| General/Stem Cell Genes | Loops disrupted | Largely unaffected | Basic cellular maintenance |
| Developmental Genes | Loops disrupted | Failed activation | Organ and tissue identity |
| Transcription Factors | Isolated/Disconnected | Impaired expression | Directing cell specialization |
Clinical Implications and Next Steps
The ability to distinguish between “resilient” and “vulnerable” genes opens new avenues for understanding how slight perturbations in DNA folding can lead to profound medical consequences. For patients with cohesinopathies or certain malignancies, the disease may not be caused by a global failure of the genome, but by the specific silencing of a few critical developmental switches.
Apostolou and her lab are now shifting their focus toward identifying exactly which genes are most dependent on cohesin and why some are more protected than others. By understanding these conditions, researchers hope to pinpoint the specific genetic triggers that, when disrupted, lead to developmental impairment or oncogenesis.
The study was supported by several major health and research organizations, including the National Institutes of Health (NIH) through the National Cancer Institute, the National Institute of Neurological Disorders and Stroke, and the Human Genome Research Institute, as well as the Tri-Institutional Stem Cell Initiative by the Starr Foundation.
Disclaimer: This article is for informational purposes only and does not constitute medical advice. Please consult a healthcare professional for questions regarding genetic disorders or cancer treatment.
The Apostolou lab will continue to analyze the interactions between architectural proteins and the genome to determine how these vulnerabilities can be mitigated or targeted in therapeutic contexts. Further updates on these genomic mapping efforts are expected as the team expands its analysis of human cell lines.
Do you have questions about the latest breakthroughs in genomic research? Share your thoughts in the comments or share this article with your network.

