The intricate process by which cells dismantle damaged or unwanted proteins, known as proteasome assembly, isn’t the rigid, step-by-step procedure scientists once believed. Modern research published in Nature and detailed by Wiley Analytical Science reveals a surprisingly flexible system with multiple pathways, challenging long-held assumptions about cellular function. This discovery, stemming from advanced structural biology techniques, could have significant implications for understanding diseases linked to proteasome dysfunction, including neurodegenerative disorders and certain cancers.
For years, the prevailing model suggested proteasomes – often described as the cell’s “recycling centers” – assembled in a highly ordered, linear fashion. Proteins would attach in a specific sequence, building the complex piece by piece. However, researchers at the Max Planck Institute of Biochemistry in Martinsried, Germany, have demonstrated that the assembly process is far more dynamic. Using cryo-electron microscopy, they observed that proteasome components can join in various orders, adapting to cellular conditions and potentially optimizing efficiency. This study in Nature provides unprecedented structural insights into these transitional states.
Unraveling the Complexity of Proteasome Assembly
Proteasomes are essential for maintaining cellular health. They degrade proteins that are misfolded, damaged, or no longer needed, preventing the accumulation of toxic substances. Dysfunction in the proteasome system is implicated in a range of diseases. For example, the buildup of misfolded proteins is a hallmark of Alzheimer’s and Parkinson’s diseases. Understanding how proteasomes are built – and what can head wrong during that process – is therefore crucial for developing effective therapies.
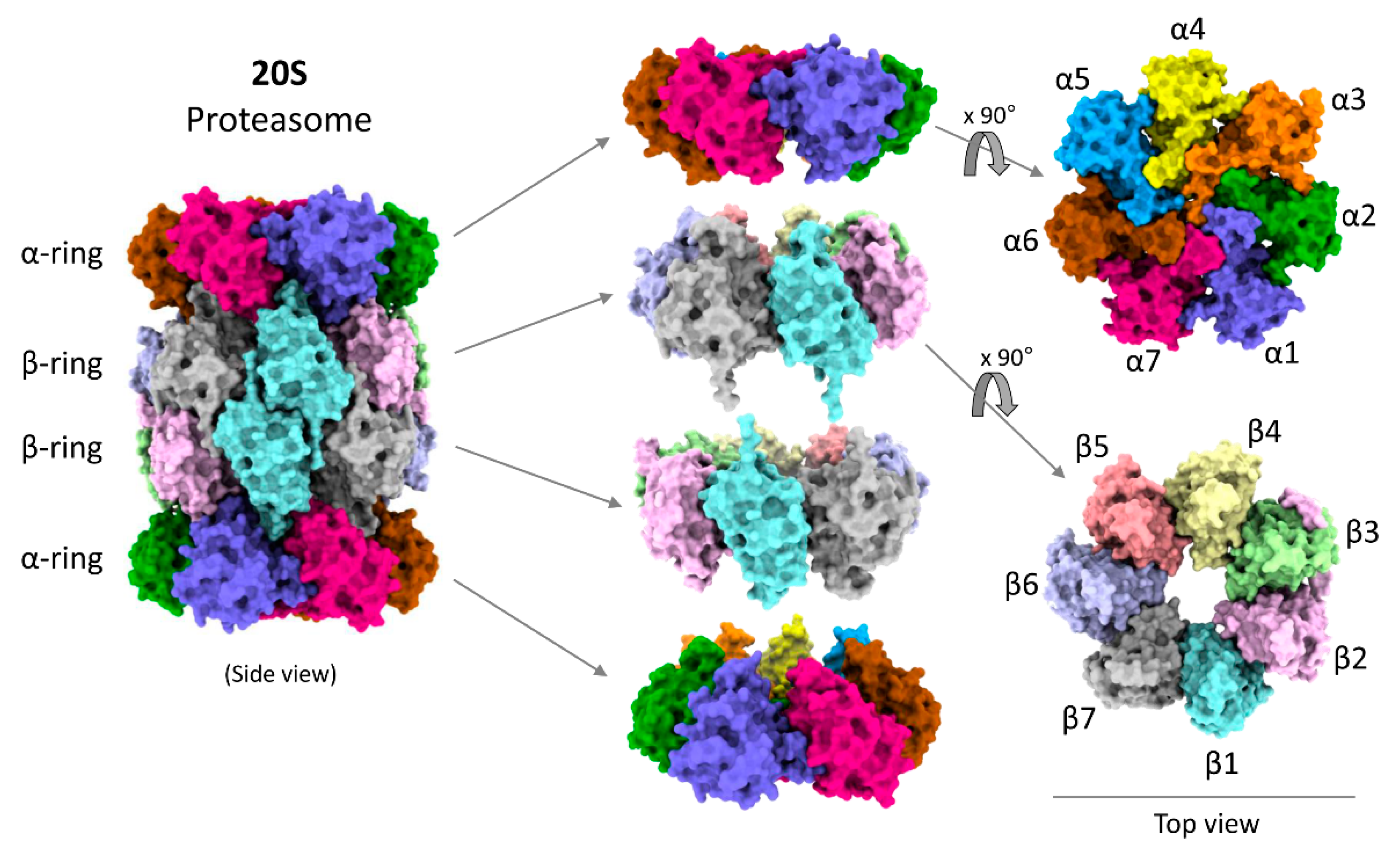
The research team, led by Dr. Wolfgang Baumeister, focused on the 20S core particle of the proteasome, the barrel-shaped structure responsible for protein degradation. They meticulously mapped the different stages of assembly, revealing that several intermediate structures exist and that components can attach and detach in a non-linear manner. “We found that the proteasome doesn’t follow a single, predetermined path,” explained Dr. Baumeister in a report from Technology Networks. “It’s more like a flexible network of possibilities.”
Multiple Pathways Offer Resilience and Adaptability
The discovery of these multiple assembly pathways suggests a built-in resilience to the system. If one pathway is blocked or compromised, the cell can utilize alternative routes to build a functional proteasome. This adaptability could be particularly important in stressful conditions or when faced with genetic mutations. Researchers believe this flexibility may similarly allow cells to fine-tune proteasome activity based on their specific needs.
The team identified at least three distinct pathways for assembling the 20S core particle. These pathways differ in the order in which specific protein subunits are added. The structural transitions observed during assembly were remarkably complex, involving significant conformational changes in the proteasome components. Cryo-electron microscopy allowed the researchers to capture these fleeting intermediate structures with unprecedented detail, providing a dynamic picture of the assembly process.
Implications for Disease Research
The implications of this research extend far beyond basic cellular biology. Many cancer cells, for instance, exhibit elevated proteasome activity, relying on the system to remove proteins that would otherwise trigger cell death. Proteasome inhibitors, such as bortezomib, are already used as chemotherapy drugs to block this activity, but cancer cells can sometimes develop resistance. Understanding the alternative assembly pathways could help researchers design new inhibitors that overcome this resistance.
Similarly, in neurodegenerative diseases, impaired proteasome function contributes to the accumulation of toxic protein aggregates. Identifying factors that disrupt proteasome assembly could lead to strategies for restoring its function and preventing disease progression. The research team is now investigating how different cellular signals and environmental factors influence the choice of assembly pathway.
Future Directions and Ongoing Research
While this study provides a significant leap forward in our understanding of proteasome assembly, many questions remain. Researchers are now working to determine how the assembly process is regulated and how it interacts with other cellular pathways. They are also exploring the possibility that different cell types may utilize different assembly pathways, reflecting their specialized functions.
Further research will focus on the role of accessory proteins that assist in proteasome assembly. These proteins, known as proteasome assembly chaperones, are thought to guide the process and ensure that the proteasome is correctly formed. The team also plans to investigate how proteasome assembly is affected by aging and disease.
The National Institutes of Health (NIH) continues to fund significant research into the ubiquitin-proteasome system, recognizing its central role in cellular health and disease. Updates on NIH-funded research can be found on their website. The findings regarding the flexible assembly of proteasomes represent a paradigm shift in the field, opening up new avenues for therapeutic intervention and a deeper understanding of the fundamental processes that govern life at the cellular level.
This research underscores the remarkable complexity and adaptability of cellular systems. As we continue to unravel these intricacies, we move closer to developing effective strategies for treating a wide range of diseases. Share your thoughts on this groundbreaking discovery in the comments below.